
Tuberculosis (TB), caused by Mycobacterium tuberculosis, remains one of the leading infectious causes of death worldwide and continues to pose a major public health challenge, particularly in low- and middle-income countries. The growing prevalence of drug-resistant forms, including multidrug-resistant (MDR-TB) and extensively drug-resistant TB (XDR-TB), further complicates disease control. These resistant strains arise from genetic mutations that reduce the effectiveness of first- and second-line drugs. While traditional diagnostic approaches have been central to TB control, some may require longer processing times or assess a limited number of resistance markers, making it difficult to comprehensively identify resistance-associated mutations and guide optimal treatment decisions.
G2M offers a rapid detection & identification platform for Mycobacterium Tuberculosis Complex and drug-resistant TB, using a targeted sequencing approach directly from clinical specimens, eliminating the need for culture.
This assay is designed to map 100 kb regions of the M. tuberculosis genome for 75 drug resistance genes and associated mutation sites, as well as SNP loci. This assay is based on Hybridization capture based target enrichment. Some of the important target genes (75) which collectively enable detection of mutations associated with resistance to first-line and second-line TB drugs include rpoB, katG,inhA, gyrA / gyrB, rrs, embB, pncA, eis etc.
With the G2M TB NGS assay, laboratories can obtain the drug-resistance profile for major first- and second-line anti-tuberculosis drugs. The panel also includes genomic regions associated with resistance to newer anti-TB agents, including the nitroimidazole antibiotic pretomanid, which is used in the WHO-recommended BPaL and BPaLM treatment regimens for drug-resistant tuberculosis. In addition, resistance-associated loci linked to other emerging drugs, including delamanid, are also covered, enabling comprehensive genomic assessment of drug resistance in Mycobacterium tuberculosis.
| First Line | Rifampicin | |
| Isoniazid | ||
| Ethambutol | ||
| Pyrazinamide | ||
| Second Line | Group A | Bedaquiline |
| Linezolid | ||
| Moxifloxacin | ||
| Levofloxacin | ||
| Group B | Clofazimine | |
| Cycloserine | ||
| Group C | Delamanid | |
| Ethionamide | ||
| Amikacin | ||
| Streptomycin | ||
| Para-aminosalicylic acid | ||
| Others | Pretomanid | |
| Capreomycin | ||
| Kanamycin |
The EZY AutoPrep-48 Automated Library Preparation Workstation streamlines NGS library preparation and improves laboratory efficiency for the TB NGS workflow.
*The panel’s performance is from the Illumina platform.
The G2M Tuberculosis Panel Reveals a Comprehensive Gene-Drug Resistance Network for Precision Therapy
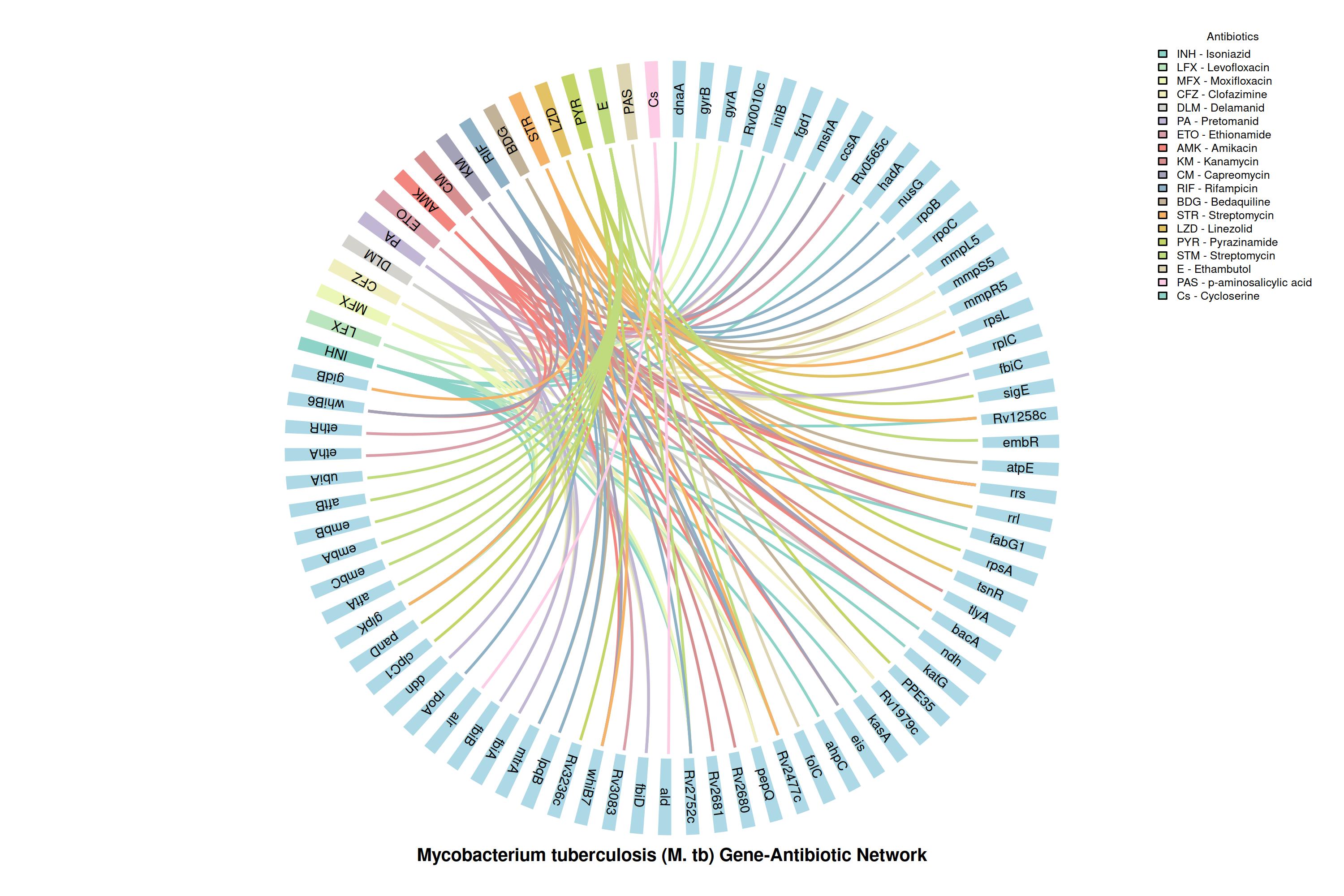
The chord diagram visualizes connections between Mycobacterium tuberculosis resistance genes and anti-TB drugs, including first line, second line, and WHO-approved regimens. This mapping highlights critical genetic determinants of MDR, Pre-XDR, and XDR-TB, enabling rapid detection and guiding precision treatment strategies by the G2M panel.
Consistent Gene Coverage and Depth Across TB Samples Confirms Assay Reliability
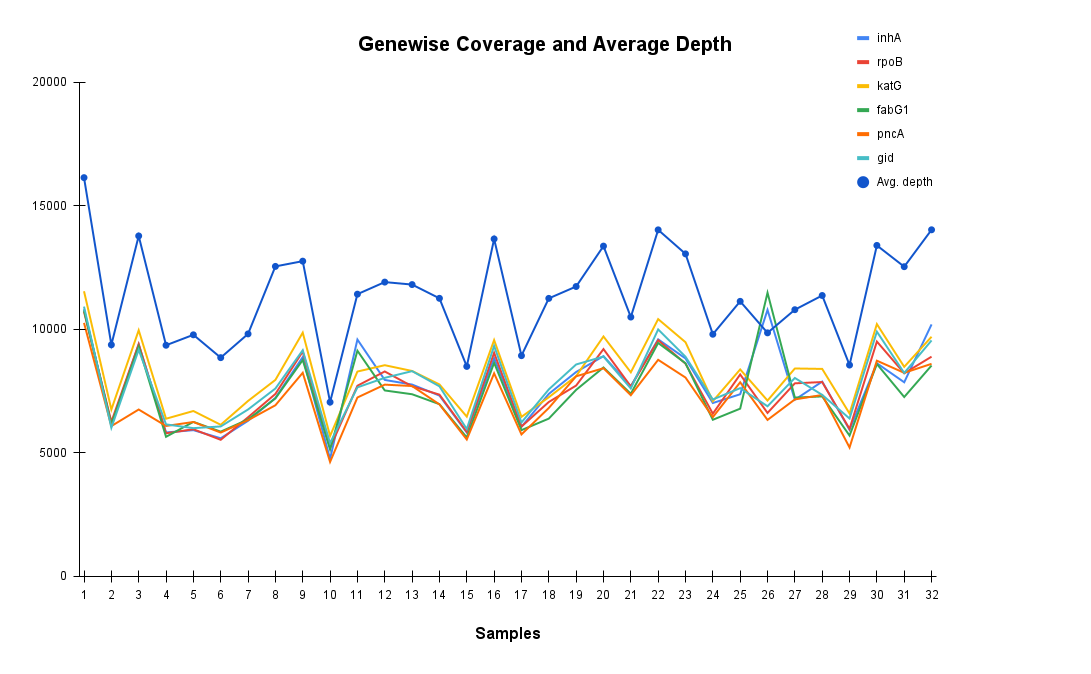
The plot illustrates gene wise coverage and average sequencing depth across 32 TB samples for critical resistance-associated genes (inhA, rpoB, katG, fabG1, pncA, gid). Coverage remains consistently high across all targets, while average depth (blue line) demonstrates robust sequencing performance. Accurate detection of resistance mutations ensures uniformity supporting reliable diagnosis and tailored therapeutic planning for MDR, Pre-XDR, and XDR-TB.
Widespread TB Drug Resistance Highlights Importance of Comprehensive Profiling
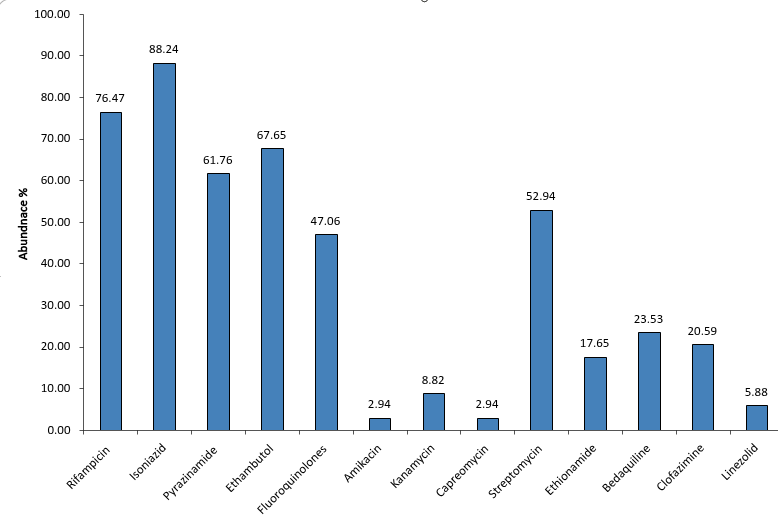
The graph illustrates the relative abundance of drug resistance across first-line and second-line anti-TB medications. Higher resistance is observed for primary drugs such as Isoniazid, Rifampicin, Pyrazinamide, and Ethambutol, followed by Fluoroquinolones. Second-line drugs including Amikacin, Kanamycin, Capreomycin, Streptomycin, Ethionamide, Bedaquiline, Clofazimine, and Linezolid show variable resistance patterns. This highlights the importance of comprehensive resistance profiling for effective TB treatment planning.
G2M TB Panel Provides Full Resistance Insight for Optimized Care
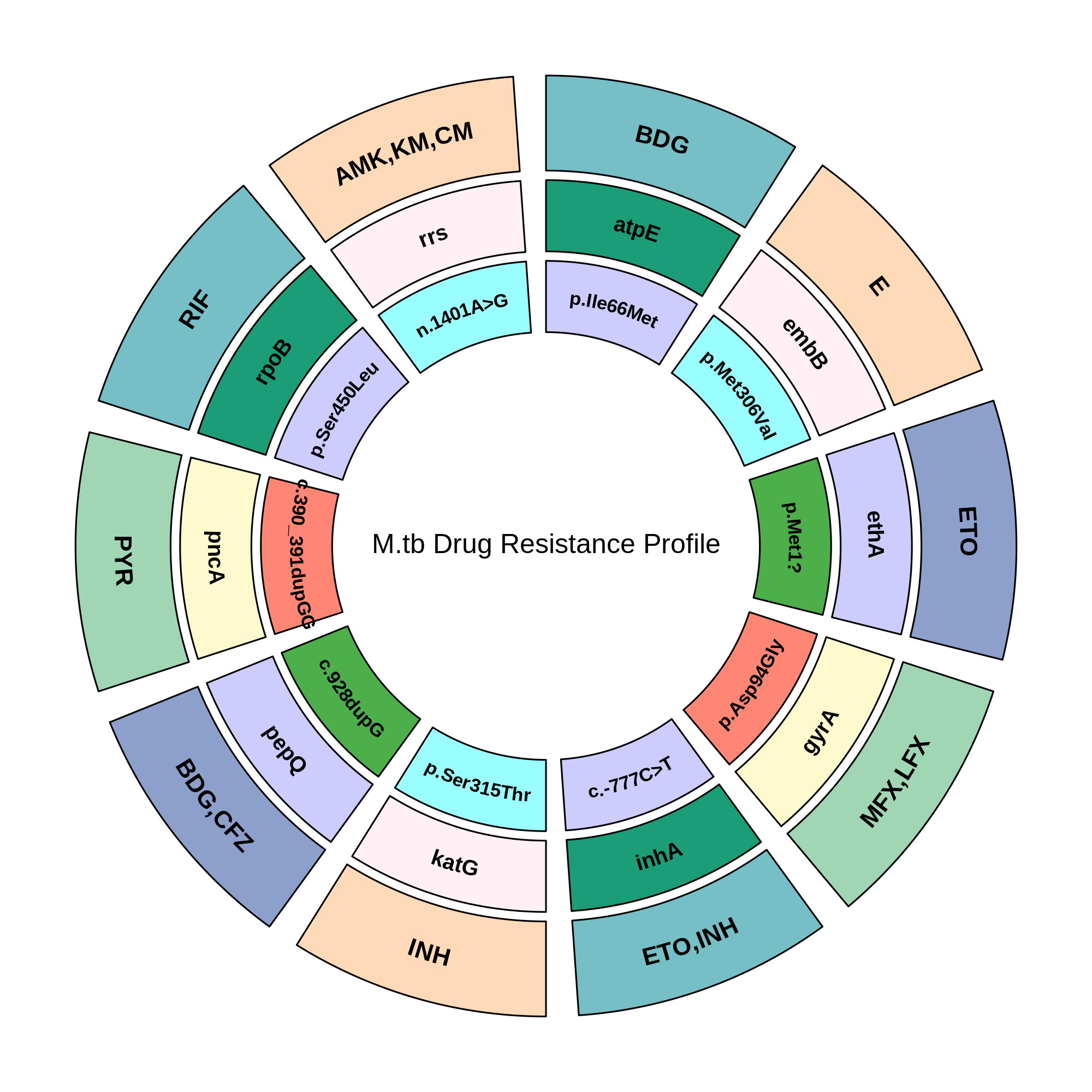
The diagram illustrates the association between Mycobacterium tuberculosis resistance genes and corresponding anti-TB drugs. Each segment represents a drug class linked to specific mutations. This integrated view highlights the genetic basis of resistance across first-line, second-line, and newer WHO recommended drugs, supporting identification to guide targeted TB therapy.
Comparison of G2M TB NGS with diffreent sample types for on-target ratio
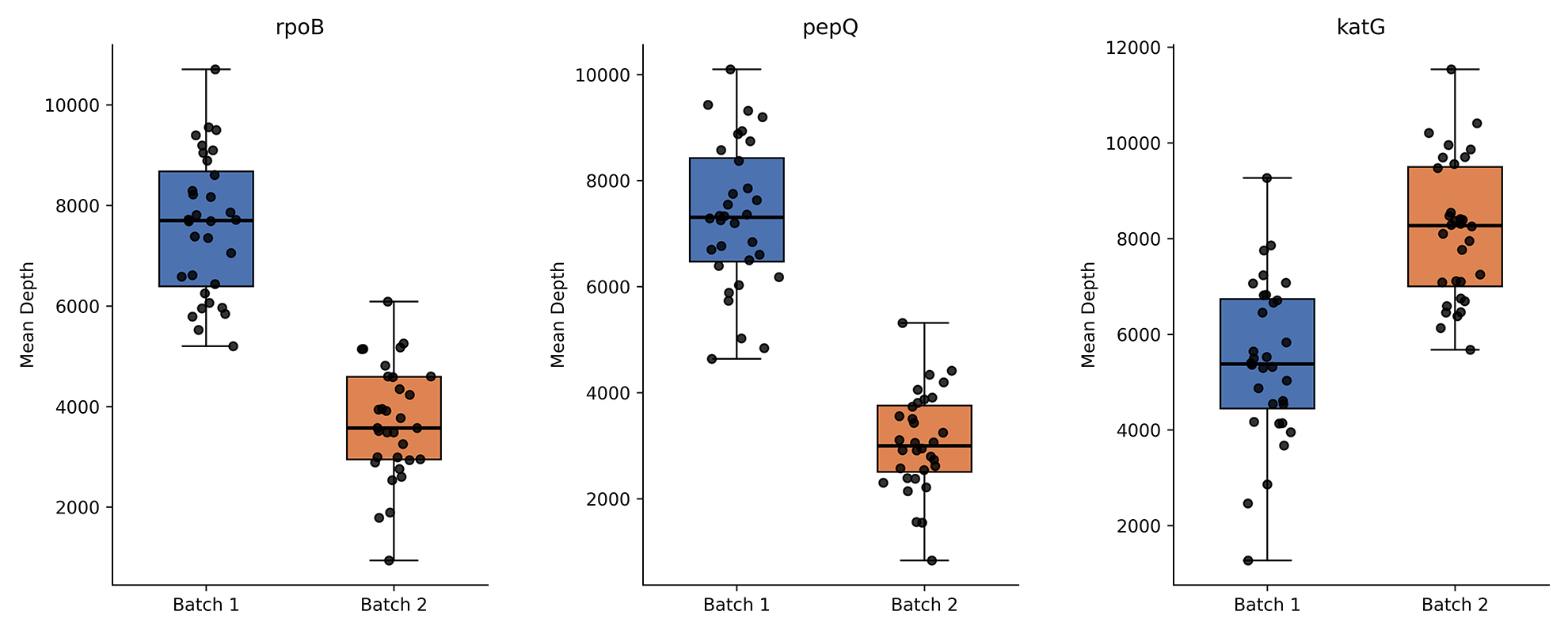
Genewise Mean depth for the G2M TB NGS assay across the clinical isolates (Batch 1) and sputum (Batch 2) samples. The bar plots demonstrate consistently and robust sequencing depth across both sample types.
High On-Target Alignment Across Different Clinical Samples
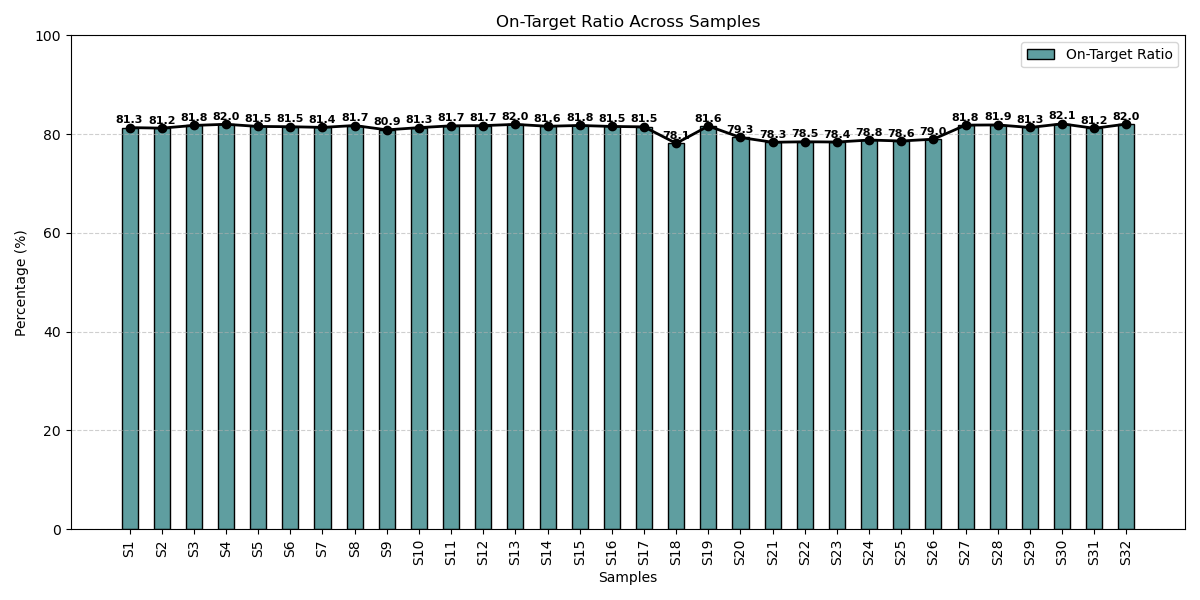
On-target ratios across TB samples consistently exceeded over 75%, highlighting the panel’s optimized design, efficient probe capture, and robust sequencing performance for reliable profiling
| Commercial Name | Cat No. | Pack Size | Platform |
|---|---|---|---|
| TB NGS test Kit | G710010-1 | 24 T | Illumina |
| G710010-2 | 96 T | Illumina | |
| G710010-3 | 96 T - EZY | Illumina - EZY | |
| TB NGS test Kit | G710010-4 | 24 T | MGI |
| G710010-5 | 96 T | MGI | |
| G710010-6 | 96 T - EZY | MGI - EZY | |
| TB NGS test Kit | G710010-7 | 24 T | Aviti |
| G710010-8 | 96 T | Aviti | |
| G710010-9 | 96 T - EZY | Aviti - EZY | |
| TB NGS test Kit | G710010-10 | 24 T | Thermo |
| G710010-11 | 96 T | Thermo | |
| G710010-12 | 96 T - EZY | Thermo - EZY |
Since its inception in 2016, Genes2me has been constantly striving towards setting a benchmark in the diagnostics space by introducing premium quality (Made in India) diagnostic kits which are CE-IVD, ISO-13485:2016, and ISO 9001:2015 certified, assuring our clients of unparalleled quality and compliance with international standards.
© 2025 Genes2me. All rights reserved.