
Liquid biopsy has emerged as a transformative approach in precision medicine, enabling non-invasive detection and monitoring of disease through analysis of circulating biomarkers such as circulating tumor DNA (ctDNA), and other nucleic acids present in blood and other body fluids. Unlike traditional tissue biopsies, liquid biopsy offers a safer, repeatable, and more comprehensive method for capturing tumor heterogeneity and real-time disease dynamics.
Next-generation sequencing (NGS) has significantly enhanced the capabilities of liquid biopsy by enabling highly sensitive and multiplexed detection of genomic alterations, including single nucleotide variants (SNVs), insertions and deletions (indels), copy number variations (CNVs), and structural rearrangements. This is particularly critical in oncology, where the fraction of tumor-derived DNA can be extremely low, requiring robust technologies that can accurately detect rare variants amidst a high background of normal DNA.
The success of liquid biopsy NGS assays depends on multiple factors, including efficient extraction of low input cfDNA, preservation of fragment integrity, minimization of technical bias, and the use of advanced library preparation chemistries and bioinformatics pipelines.
The Genes2Me Liquid Biopsy portfolio offers targeted, high-performance NGS panels for lung, breast, and colorectal cancers, enabling comprehensive profiling of key cancer-associated genes. These assays are optimized to work with low-input cfDNA (10–20 ng) while maintaining high sensitivity for detecting SNVs, InDels, CNVs, and gene fusions, even at low variant allele frequencies.
High sensitivity, uniform genome coverage, and error suppression strategies are essential to ensure reliable detection, especially in challenging samples with low variant allele frequencies.
Built with a focus on efficient cfDNA extraction compatibility, robust library preparation, and uniform genome coverage, our panels ensure reliable and reproducible results. High-depth sequencing (≥20,000×) and optimized target capture enable strong on-target performance and consistent coverage across clinically relevant regions, minimizing technical bias and enhancing detection confidence.
From sample to report, the Genes2Me liquid biopsy solutions support laboratories with a scalable and integrated workflow—empowering high-throughput testing while maintaining accuracy and operational efficiency. Whether for focused panels or comprehensive genomic profiling, our solutions are designed to deliver clinically actionable insights with precision and consistency.

The RapiX-LB is an advanced, fully automated system specifically designed for the extraction of circulating DNA from liquid biopsy samples. Engineered to handle high sample input volumes of up to 3 mL, it ensures efficient recovery of low-abundance nucleic acids critical for downstream molecular applications. Capable of processing up to four samples simultaneously, the RapiX-LB utilizes prefilled reagent cartridges for maximum user convenience and minimal hands-on time.

The workflow is designed to be automation-friendly, supporting both manual and automated library preparation formats to accommodate varying laboratory throughput requirements. The Assay demonstrates platform-agnostic compatibility and has been validated across commonly used sequencing systems, including Illumina, Element Biosciences, MGI, and Thermo Fisher platforms.
NGS data analysis is supported by GATK-based pipelines integrated with the Cliseq Interpreter Platform, a cloud-based clinical interpretation solution designed to streamline analysis of complex genomic data.
*The panel’s performance is from the Illumina platform.
High-Fidelity Coverage for Confident Variant Detection
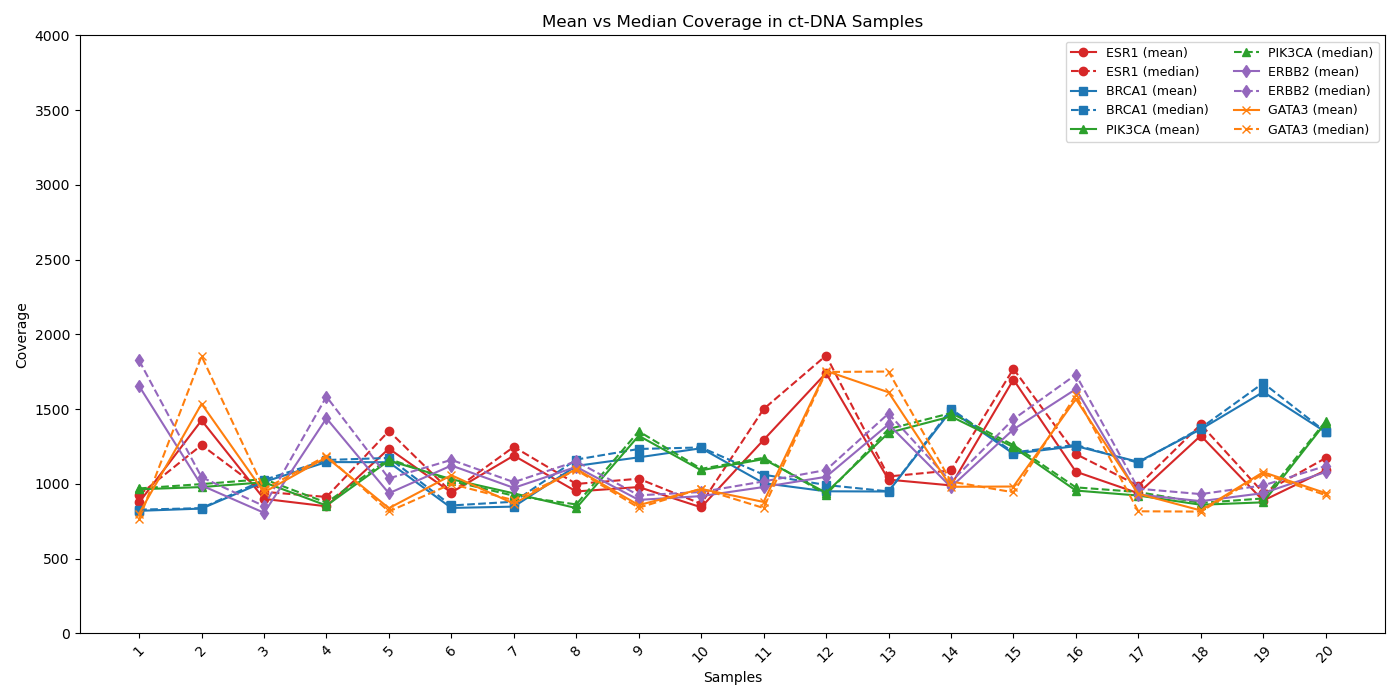
Coverage profiles of critical genes (ESR1, BRCA1, PIK3CA, ERBB2, and GATA3) exhibit strong concordance between mean (solid line) and median (dashed line) depth, demonstrating uniform sequencing and minimal bias across all target regions. This alignment highlights the assay’s robust performance and reliability, ensuring confident results across diverse breast cancer samples.
Mutation Burden Landscape Across Breast Cancer Genes
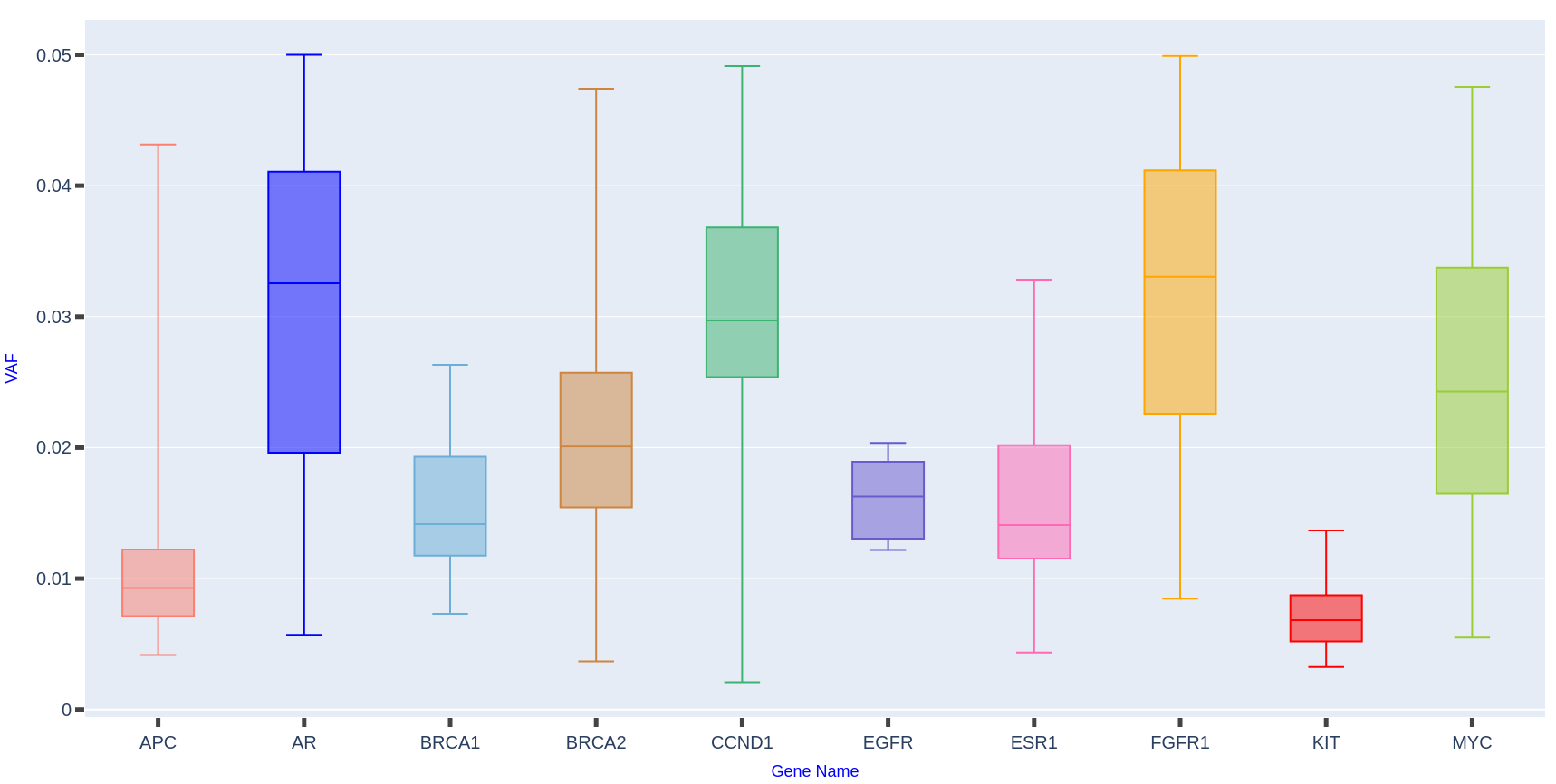
Distribution of variant allele frequencies (VAFs) in 158 breast cancer samples reveals distinct mutation patterns. Genes such as AR, CCND1, and FGFR1 exhibit higher median VAFs with broad variability, indicating a substantial mutation burden and potential influence on disease progression. Conversely, APC, EGFR, and KIT show lower, more uniform VAFs, suggesting a stable or limited role in the overall genomic architecture.
Efficient On-Target Capture for Liquid Biopsy Precision
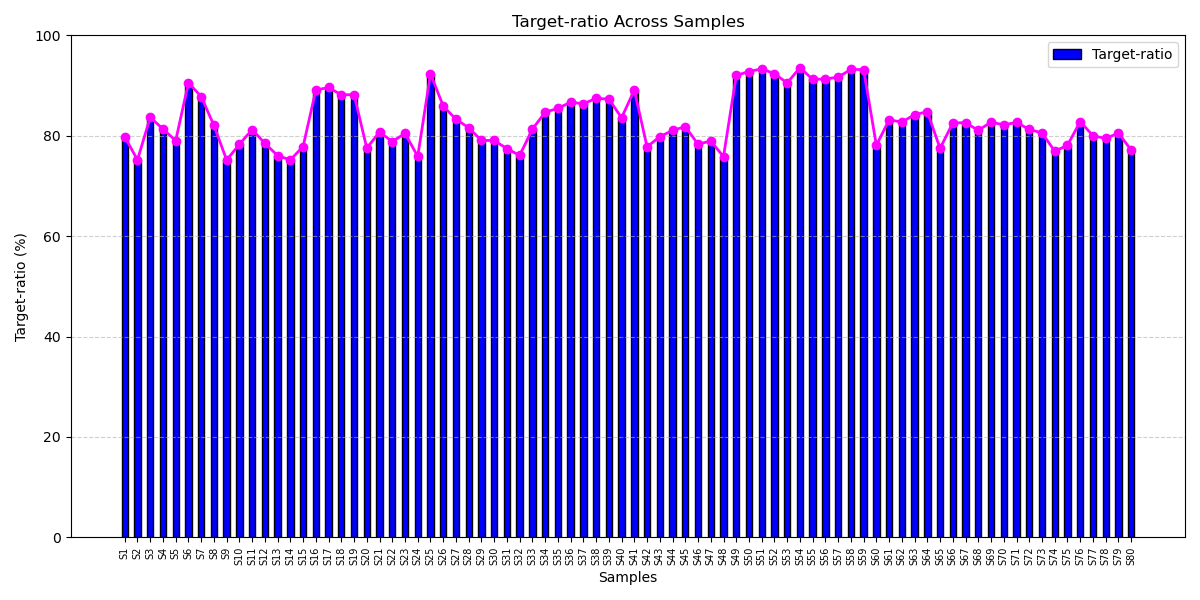
The ctDNA breast cancer panel consistently delivers more than 75% on-target alignment, reflecting its smart design and efficient target capture. This ensures reliable mutation detection from low-input samples, enabling early diagnosis, treatment monitoring, and scalable diagnostic utility.
Uniform BRCA1 Exon Coverage with G2M ctDNA Breast Cancer Panel
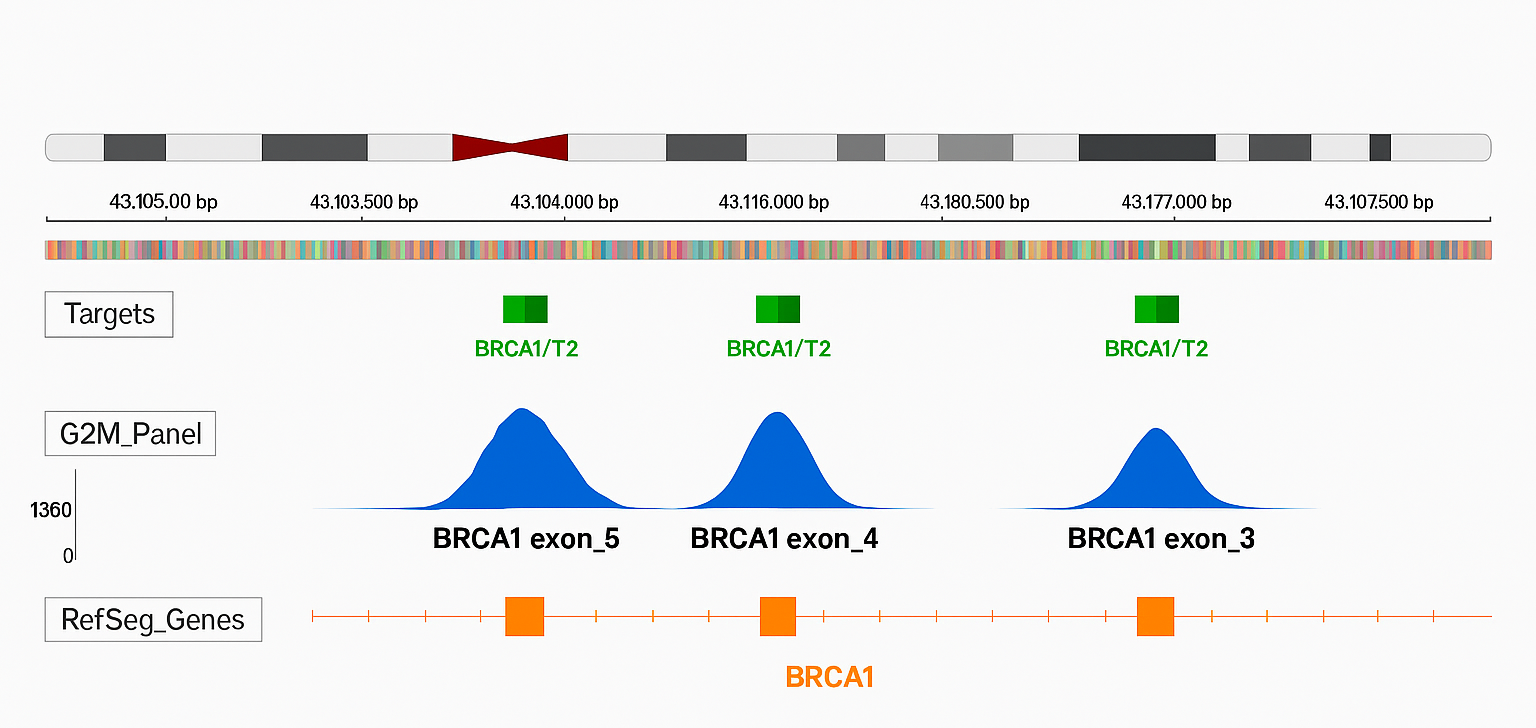
The figure illustrates read coverage for BRCA1 exons 3, 4, and 5. The top panel highlights the BRCA1 target region (green), the middle panel displays per-base coverage depth achieved by the G2M ctDNA Breast Cancer Panel (blue), and the bottom panel represents the gene’s coding regions as annotated by RefSeq (orange). This visualization underscores uniform coverage across clinically significant exons, ensuring reliable variant detection.
Mutation Hotspots Driving Breast Cancer Genomics
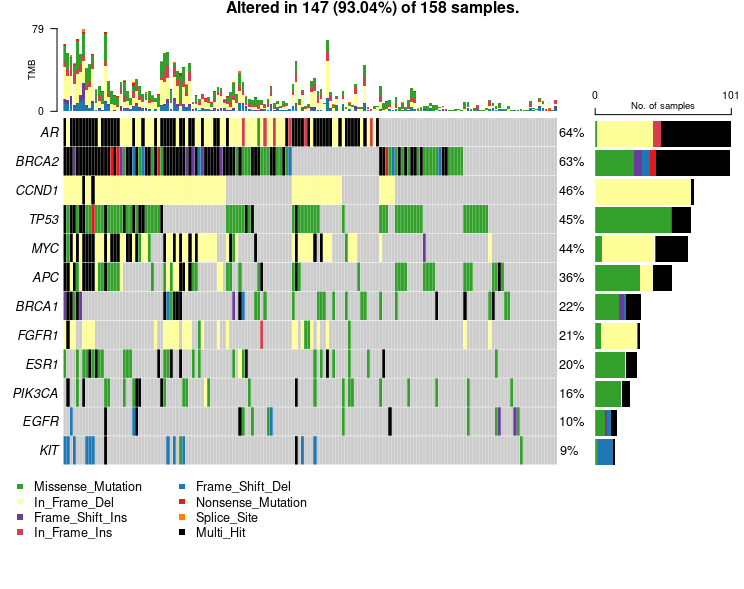
Oncoplot showcasing the 12 most frequently mutated genes in ctDNA breast cancer samples. Missense mutations (green) dominate, followed by in-frame deletions (yellow), while multi-hit events (black) indicate multiple mutation types within the same gene. The right panel summarizes mutation frequency per gene.
| Commercial Name | Cat No. | Pack Size | Platform |
|---|---|---|---|
| ctDNA Lung NGS Test Kit | G710028-1 | 24 T | Illumina |
| G710028-2 | 96 T | Illumina | |
| G710028-3 | 96 T – EZY | Illumina – EZY | |
| G710028-4 | 24 T | MGI | |
| G710028-5 | 96 T | MGI | |
| G710028-6 | 96 T – EZY | MGI – EZY | |
| G710028-7 | 24 T | Aviti | |
| G710028-8 | 96 T | Aviti | |
| G710028-9 | 96 T – EZY | Aviti – EZY | |
| G710028-10 | 24 T | Thermo | |
| G710028-11 | 96 T | Thermo | |
| G710028-12 | 96 T – EZY | Thermo – EZY | |
| ctDNA Colorectal NGS Test Kit | G710029-1 | 24 T | Illumina |
| G710029-2 | 96 T | Illumina | |
| G710029-3 | 96 T – EZY | Illumina – EZY | |
| G710029-4 | 24 T | MGI | |
| G710029-5 | 96 T | MGI | |
| G710029-6 | 96 T – EZY | MGI – EZY | |
| G710029-7 | 24 T | Aviti | |
| G710029-8 | 96 T | Aviti | |
| G710029-9 | 96 T – EZY | Aviti – EZY | |
| G710029-10 | 24 T | Thermo | |
| G710029-11 | 96 T | Thermo | |
| G710029-12 | 96 T – EZY | Thermo – EZY | |
| ctDNA-Breast NGS Test Kit | G710011-1 | 24 T | Illumina |
| G710011-1-2 | 96 T | Illumina | |
| G710011-1-3 | 96 T – EZY | Illumina – EZY | |
| G710011-1-4 | 24 T | MGI | |
| G710011-1-5 | 96 T | MGI | |
| G710011-1-6 | 96 T – EZY | MGI – EZY | |
| G710011-1-7 | 24 T | Aviti | |
| G710011-1-8 | 96 T | Aviti | |
| G710011-1-9 | 96 T – EZY | Aviti – EZY | |
| G710011-1-10 | 24 T | Thermo | |
| G710011-1-11 | 96 T | Thermo | |
| G710011-1-12 | 96 T – EZY | Thermo – EZY |
Since its inception in 2016, Genes2me has been constantly striving towards setting a benchmark in the diagnostics space by introducing premium quality (Made in India) diagnostic kits which are CE-IVD, ISO-13485:2016, and ISO 9001:2015 certified, assuring our clients of unparalleled quality and compliance with international standards.
© 2025 Genes2me. All rights reserved.