From Direct Blood to Results in ~2 Hours: A Comprehensive RT-PCR Approach to Culture-free Sepsis and AMR Detection

Sepsis and AMR is one of the most urgent and complex challenges in modern medicine. Despite advances in critical care and infectious disease management, it continues to claim millions of lives each year. The World Health Organization (WHO) estimates that sepsis affects about 49 million people around the world and causes about 11 million deaths each year, which is almost one in five deaths worldwide. Every hour that goes by without effective antimicrobial therapy, the risk of death goes up a lot.
At the same time, the crisis is getting worse because of antimicrobial resistance (AMR). The Center for Disease Control and Prevention (CDC) estimates that in the United States alone, more than 2.8 million antimicrobial-resistant infections occur annually, leading to over 35,000 deaths. Globally, AMR is projected to cause up to 10 million deaths per year by 2050, if left unchecked.
The intersection of Sepsis and Antimicrobial resistance (AMR) presents a diagnostic paradox. Healthcare professionals (HCP’s) must start rapid, broad-spectrum therapy right away to save lives, but using antibiotics incorrectly makes resistance worse. Therefore, the need for faster, more comprehensive diagnostics has never been more pressing.
The Broader Context: Sepsis and AMR as Converging Global Threats
The clinical overlap between sepsis and AMR is real and significant. Sepsis is often treated empirically with broad-spectrum antibiotics due to the lack of rapid pathogen identification. While necessary, this approach can inadvertently promote antimicrobial resistance (AMR). Conversely, infections caused by resistant pathogens are more likely to progress to sepsis when first-line therapies fail to control the infection promptly.
Bloodstream infections(BSIs), a major cause of sepsis, frequently involve bacteria with resistance traits that complicate effective antibiotic therapy. These resistant infections are associated with higher mortality, prolonged illness, and longer ICU stays compared with susceptible infections.
Therefore, the urgency of integrated diagnostics is underscored by the growing overlap between sepsis and multidrug-resistant organisms. Carbapenem-resistant Enterobacterales, methicillin-resistant Staphylococcus aureus, and other resistant pathogens are increasingly implicated in bloodstream infections. In many regions, AMR rates exceed 30–50% for key pathogens. As resistance spreads, empiric therapy becomes less reliable. The traditional “start broad and wait” strategy becomes riskier and more resource intensive.
Molecular assays that provide rapid organism identification along with genotypic resistance information within hours offer a critical diagnostic strategy to address this growing clinical challenge.
The Diagnostic Bottleneck in Sepsis Care
The gold standard for sepsis diagnosis remains blood culture, followed by phenotypic Antimicrobial Susceptibility Testing (AST). While culture-based workflows are highly specific, they are inherently time-consuming:
- Blood culture positivity: 12–48 hours
- Pathogen identification: additional hours
- Phenotypic AST: 24–72 hours post-isolation
In practice, doctors usually wait 48 to 96 hours to get useful resistance data. During this time, patients might get broad-spectrum antibiotics that aren’t always needed or aren’t enough. Moreover, blood cultures have well-documented limitations:
- Less sensitivity in patients who have already been treated with antibiotics
- Low levels of pathogens in the early stages of sepsis
- Difficulty detecting fastidious or slow-growing organisms
Molecular diagnostics have helped bridge some of these gaps by making it possible to identify organisms more quickly and, in some cases, find specific resistance markers. But a lot of the assays that are available now are still fragmented. For example, they need separate panels to detect pathogens and AMR genes, or they need positive blood cultures as input.
This fragmentation makes workflows more complicated, takes longer to turn around, and puts more work on laboratories.
Introducing a New Approach: Combining Pathogen and Resistance Detection in One
To address this unmet clinical need, G2M has developed SepsiAMR-Q, an advanced, all-in-one RT-PCR solution designed for simultaneous detection of key sepsis-causing pathogens and critical antimicrobial resistance (AMR) genes. The SepsiAMR-Q assay was developed with a simple premise: in sepsis management, pathogen identity and resistance profile are inseparable clinical questions.
Rather than running separate panels or reflex tests, this comprehensive multiplex assay interrogates:
- Major Gram-positive pathogens
- Major Gram-negative pathogens
- Fungal targets relevant to bloodstream infections
- Critical AMR genes, including those conferring resistance to β-lactams, carbapenems, and other high-priority antibiotic classes
This assay utilizes advanced real-time PCR technology for the qualitative and parallel detection of 32 sepsis-causing pathogens and 18 antimicrobial resistance (AMR) genes directly from human blood culture and whole blood samples and includes an internal control in each tube to ensure assay integrity. With ready-to-use lyophilized reagents and an 8-sample assay format, SepsiAMR-Q offers a reliable and efficient approach for rapid sepsis diagnosis and resistance profiling.
This dual-detection approach addresses an important clinical reality: identifying both the key genes and the pathogens significantly influences therapeutic decision-making. For example, knowing that a patient has Klebsiella pneumoniae, is useful, but knowing it harbours a carbapenemase gene such as KPC or NDM fundamentally changes therapeutic decisions.
The Direct Blood Advantage
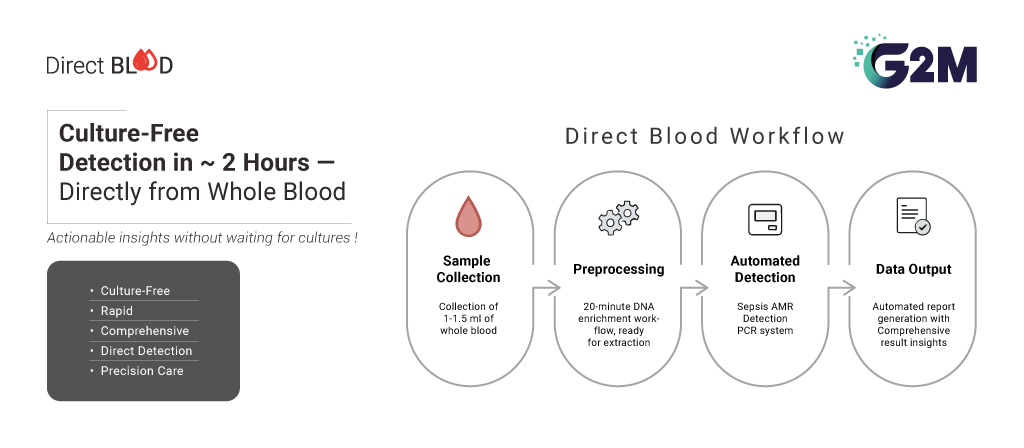
Perhaps the most transformative element of this approach is its ability to work directly from whole blood, as well as from positive blood cultures. Traditional workflows depend on microbial amplification via culture to achieve detectable concentrations. However, molecular detection from whole blood eliminates the culture step entirely, enabling culture-free identification in less than two hours.
The implications are significant:
- Reduced time to result from days to hours
- Improved detection in patients already receiving antibiotics
- Earlier antimicrobial optimization
- Potential reduction in ICU length of stay
Early data from rapid molecular diagnostics consistently demonstrate that shortening time to targeted therapy correlates with improved clinical outcomes and reduced mortality risk. Rapid exclusion of resistant determinants can support early de-escalation from broad-spectrum therapy, minimizing unnecessary antibiotic exposure.
A Closed, End-to-End Workflow
Sepsis diagnostics must operate reliably under high-pressure clinical conditions. Open workflows increase contamination risk, require multiple instruments, and demand significant technical oversight.
The SepsiAMR-Q platform addresses these challenges through a fully integrated, closed system architecture- the SepsiAMR PCR System capable of both extraction and amplification.
This end-to-end configuration minimizes manual intervention, reduces cross-contamination risk, and standardizes performance across runs.
High Throughput Without Compromising Speed
In many institutions, sepsis testing volume is substantial, particularly in emergency departments and intensive care units. Rapid tests that handle only a few samples at a time can create bottlenecks.
The SepsiAMR-Q workflow achieves an 8-sample throughput per run while maintaining rapid turnaround. Combined with its extraction system, laboratories can process multiple samples efficiently without sacrificing the sub-two-hour time frame for direct blood detection.
This combination of speed and throughput addresses a common tension in molecular diagnostics which is balancing urgent clinical timelines with operational realities.

Clinical and Public health Implications
The integration of comprehensive pathogen and AMR gene detection into a single workflow has implications beyond individual patient management.
1. Improved Clinical Outcomes-Rapid identification of resistant organisms supports timely escalation to appropriate therapy, which is strongly associated with reduced mortality in septic shock.
2. Antimicrobial Management-Early differentiation between resistant and susceptible infections can reduce unnecessary carbapenem or last-line antibiotic use.
3. Infection Control-Prompt detection of high-risk resistance determinants (e.g., carbapenemases) enables faster isolation decisions and outbreak mitigation.4. Economic Impact-Sepsis is one of the most expensive conditions treated in hospitals worldwide. Shortening ICU stays and reducing broad-spectrum antibiotic usage may yield meaningful cost savings.
Moving from Reactive to Proactive Sepsis Management
Sepsis care has historically been reactive; waiting for cultures, escalating therapy upon deterioration, and adjusting treatment when susceptibility results arrive. The next phase of diagnostics aims to shift this paradigm toward proactive, precision-guided intervention at the earliest possible stage.
Comprehensive, closed-system RT-PCR platforms such as SepsiAMR-Q reflect this evolution. By unifying pathogen and AMR detection, enabling direct-from-blood analysis, and reducing workflow complexity, these systems illustrate how molecular diagnostics can reshape critical care microbiology.
